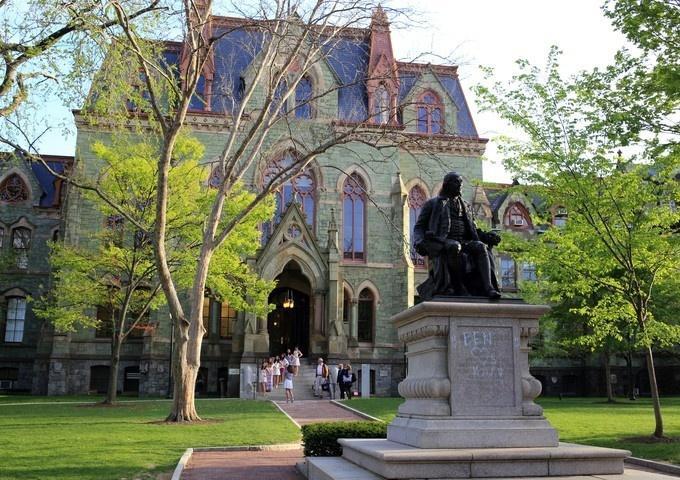

宾夕法尼亚大学(宾大,University of Pennsylvania ),于1740年始建,位于美国宾夕法尼亚州费城,是一所全球顶尖的私立研究型大学,著名的八所常春藤盟校之一,北美顶尖大学学术联盟美国大学协会14所创始成员之一。
想要出国访问学习的小伙伴看过来啦,美国宾夕法尼亚大学生物学、免疫学、生物化学或计算机科学、工程、生物信息学方向正在招收访问学者、博士后!51访学网小编每周定时更新最新的访学招聘信息,感谢关注51访学。
Fellow positions at Penn Epigenetics Institute: Philadelphia, PA, United States.
Positions for wet (experimental), dry (computational), or hybrid biologists are available in the interdisciplinary lab of Dr. Golnaz Vahedi.
The Vahedi laboratory is part of the Department of Genetics, Perelman School of Medicine at the University of Pennsylvania.
We are a core member of the Epigenetics Institute (https://goo.gl/GnaZdx) and a member of the Institute for Immunology (https://goo.gl/Vi2jAP).
We are a member of the Human Pancreas Analysis Program at Penn (https://hpap.pmacs.upenn.edu/) and have an unprecedented opportunity to perform deep profiling of the human endocrine pancreas and its interaction with the immune system.
Our laboratory is multidisciplinary, integrating computational and experimental approaches to develop a system-level understanding of gene regulation in immune cells.
We blend epigenomics, human and mouse genetics, immunology, and computational biology to pursue a new understanding of gene regulation in the immune system.
We use bulk assays including ChIP-seq, ATAC-seq, and chromatin conformation assays such as HiC, in addition to cutting-edge single-cell genomic assays such as single-cell (sc)ATAC-seq and scRNA-seq or the single-cell Oligopaint imaging technique.
Our lab’s major contributions in less than 5 years :
(1) the discovery of genome misfolding in type 1 diabetes (https://www.cell.com/immunity/fulltext/S1074-7613(20)30030-3 received major media attention and featured on journal’s cover).
(2) the discovery of TCF-1 as a pioneer factor in T cells (https://www.cell.com/immunity/fulltext/S1074-7613(18)30033-5 featured on journal’s cover).
(3) the development of a novel computational method for the joint profiling of chromatin accessibility and CAR-T integration site analysis at the single-cell level (https://www.pnas.org/content/early/2020/02/19/1919259117.short)
The lab is funded by National Institutes of Health. Our contribution to single-cell genomic field was recently recognized by a Chan Zuckerberg Initiative award.
Experimental candidates are required to hold a PhD or a visiting scholar in molecular biology, immunology, or biochemistry. Experience in genomics and epigenomics will be viewed favorably.
Potential training opportunities exist for the experimental scientist joining our lab to learn computational genomics.
Computational candidates are required to hold a PhD in computer science, engineering, bioinformatics or related programs. Expertise in machine learning and statistics in addition to programing in Python and R in a Unix environment is required. Trainees interested in image processing of 3D FISH imaging data are encouraged to apply.
Further information about our lab could be found at http://vahedilab.github.io/LabSite/
Interested applicants should submit a CV to vahedilab.jobs@gmail.com.
51访学网专注国外访问学者申请服务,已积累不计其数的成功医学访学案例,如安德森癌症中心、梅奥诊所、麻省总医院、布莱根妇女医院、斯坦福医院、JHU医院、克利夫兰诊所等顶尖医学中心访学案例举不胜举。 因此,如果你对访学有所兴趣不妨试一下。希望我们能实现您医学访学的梦想!更多医生访学相关问题,请添加51访学专业咨询顾问于老师微信:woyaofangxue 想了解更多请点击阅读原文,留下您的联系方式,我们专业咨询顾问会和您联系~ 51访学网精彩文章点击查看↓ ↓ ↓ ↓ 51访学网访学案例点击查看 ↓ ↓ ↓ ↓ 霍普金斯医院| 范德堡大学 | 克利夫兰 | 麻省大学波士顿分校 | 佛罗里达大学 | 密苏里大学 | 耶鲁大学 | 匹兹堡大学医学院 | 普渡大学 | 芝加哥大学 | 哈佛大学麻省总医院 | 剑桥大学 | 麻省理工学院2020年国家留学基金资助出国留学人员选派简章出炉!
申请医学访问学者有哪些需要注意的?
医生访学需要准备什么材料?什么手续?
CSC公派申请教程-史上最全!(附资料分享)
CSC访问学者公派出国手续指南
CSC公派访问学者申请表如何填写?
如何申请及填写DS2019、DS160表!
国家选派联培博士,相关材料准备好了吗?














